BioSakshat: Training modules
Programming in R
Basic R for Life Sciences
|
R is one of the widely used popular programming languages for biological data processing and analysis. Attending Basic R for Life Sciences course will enable you to learn basic concepts of R along with hands-on experience on solving biological problems. In this training you will be introduced with its applicability to real-time problems.
|
Advanced R
|
This course is in continuation to Basic R for Life Sciences course or if you know R and you want to advance your knowledge. It introduces you to advanced concepts of R programming, which will help you to develop robust, powerful and optimized software. Topics include data manipulation using packages such as dplyr, data.table, reshape2 and others, which are very handy in terms of underlying syntax, logic and also powerful and efficient in terms of their computation. The course will cover concepts of parallel processing and strategies to optimize your code in terms of speed, exception handling, debugging and object oriented programming.
|
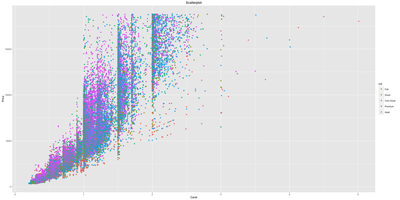
Data Visualization using R
|
A dedicated course on Data Visualization in R. The ggplot2 is one of the widely used data visualization framework developed in R. It has it's own grammar and underlying structure to it. The base graphics plotting is very different from ggplot2 plotting. Once you understand the basic concepts and logic of plotting behind ggplot2, you will be amazed how easy it is to use ggplot2 to make better quality scientific data visualizations. Apart from this, the course includes other widely used visualization packages.
|
Statistical Data Analysis using R
|
A dedicated course on Statistical Data Analysis using R. In this course, we will walk you through the fundamental statistical concepts and their application for biological data analysis with the help of R. It includes descriptive statistics, inferential statistics, hypothesis testing, regression, clustering, multivariate analysis etc.
|
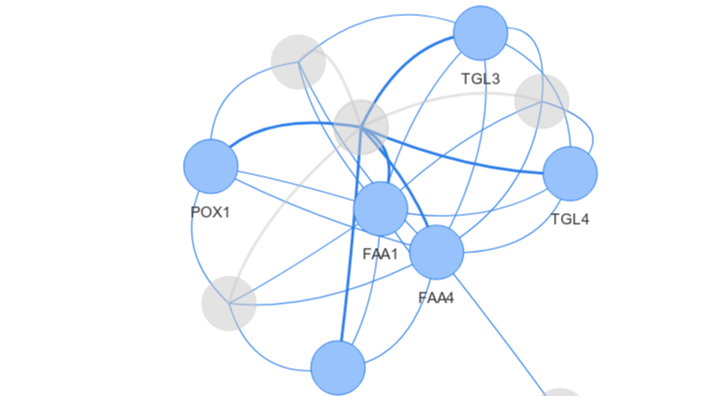
Network Analysis using R
|
A dedicated course for Social/Biological Network Analysis using R. This course focuses on Network analysis and visualization in R. Course focuses on creating networks from nodes and edges, explores various network analysis such as centrality, clique detection, clustering etc, illustrates various network visualization plots, topologies, both static and interactive plot. You will be introduced to widely used R packages for network analysis. Includes Biological network analysis with case study.
|
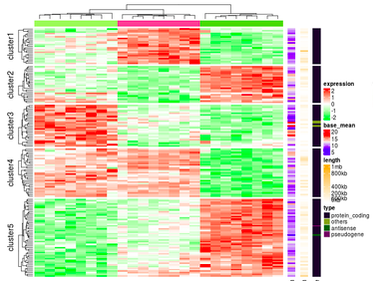
Simple to Complex Heatmaps in R
|
A dedicated course for mastering heatmaps in R. This course aims to train you on drawing heatmaps using R. Course covers simple to advanced heatmap drawing and teach you different tuning parameters to customize heatmaps. Enroll for this mini course on heatmaps.
|
Programming in Perl
Basic Perl for Life Sciences
|
Perl is one of the widely used popular programming languages for biological data processing and analysis. The knowledge of Perl will greatly help you to meet the everyday challenges posed by your research data and can save a lot of your time and efforts to deal with it. Attending Basic Perl for Life Sciences course will enable you to learn basic concepts of Perl along with hands-on experience on solving biological problems. In this training you will be introduced with its applicability to real-time problems.
|
Programming in Python
Python for Data Analysis
|
A dedicated course on Python. Python is becoming more and more popular to import, pre-process, analyse and visualize data. Python has become one of the favorite tool of choice for data science. Researchers, data analysts and software developers worldwide are using Python to harvest insights from their data in reproducible manner. Unlike any other Python course, this course focuses on Python specifically for data science enthusiasts and Life Science students and researchers.
|
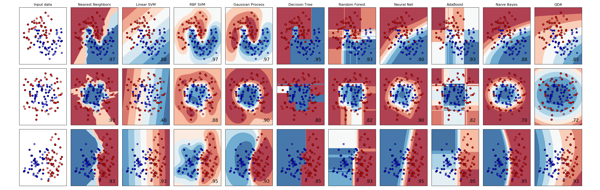
Machine Learning
|
This course is dedicated for Machine Learning using Python scikit learn.
|
Next Generation Sequencing
Whole Exome Sequencing / DNAseq data Analysis
|
A dedicated course for DNAseq data analysis. DNAseq / Whole Genome or Exome sequeincing data analysis is one of the widely practiced exercise to study somatic and germline variants in biological systems. The knowledge of WES/WGS data analysis pipeline will greatly help you to understand the details of variant calling methods used to detect somatic and germline mutations and downstream annotation methods. Attending Exome sequencing Data Analysis course will enable you to learn basic concepts of Whole genome/exome along with hands-on experience on solving real biological data.
|
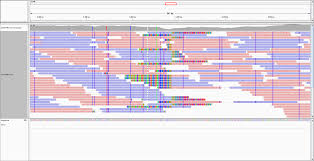
Transcriptome / RNASeq data analysis
|
A dedicated course on RNAseq data Analysis. Transcriptome / RNASeq data analysis is one of the widely practiced exercise to study differential gene expression in biological systems. Attending RNASeq Data Analysis course will enable you to learn basic concepts of RNASeq along with hands-on experience on solving real biological data.
|
ChIP-Seq Data Analysis
|
A dedicated course on ChIP-Seq data analysis. Chromatin Immunoprecipitation / ChIP-Seq data analysis is one of the widely practiced exercise to study differential binding of proteins under different conditions in biological systems. Chip-seq holds many promises for studying gene regulation, such as identification of in vivo transcription factor binding sites, histone modifications etc. Attending ChIP-Seq Data Analysis course will enable you to learn basic concepts of ChIP-Seq along with hands-on experience on solving real biological data.
|
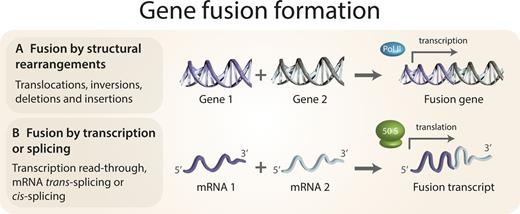
Gene Fusion
|
A dedicated course on understanding Gene Fusion and its detection using RNAseq data. Gene fusion is a chromosomal rearrangement event which plays a significant role in cancer due to the oncogenic potential of the chimeric protein generated through fusions. This module is designed to enable you to understnd end to end pipeline employed for fusion gene detection, annotation and visualization.
|
Multi-omics Data Analysis
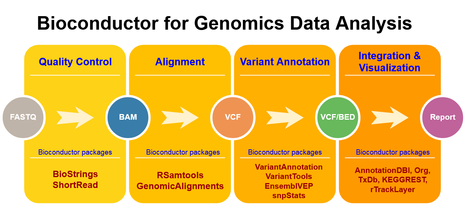
Genomics Data Analysis using Bioconductor
|
Objective of this course is to introduce you to Bioconductor for analysis of NGS based genomics data. Bioconductor repository contains several R packages that allow to perform rigorous statistical analyses and visualization of large-scale omics data. In this course we will teach you Bioconductor by taking whole genome/exome sequencing (WGS/WES) data analysis as case study.
|
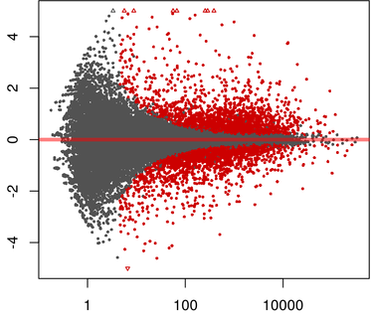
Multi-Transcriptomics Data Analysis using Bioconductor
|
Objective of this course is to introduce you to the general practices for transcriptome data analysis using advanced statistical models. We will use DESeq2 / edgeR / Limma, Bioconductor packages which are widely used for processing RNASeq read count data. Theses packages support count-based statistical models that expects input data in the form of matrix of raw read counts. This read count matrix after normalization is an approximation of transcript abundance.
|
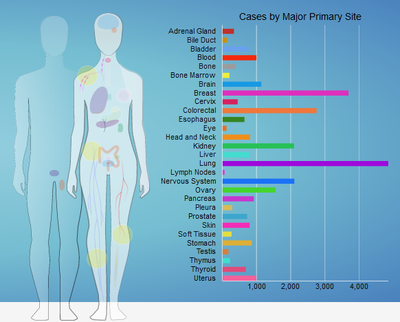
Analysis of multi-omics cancer cohort data from TCGA
|
This course introduces you to analyze multi omics data from TCGA. Please click on know more to glance through the course content.
|
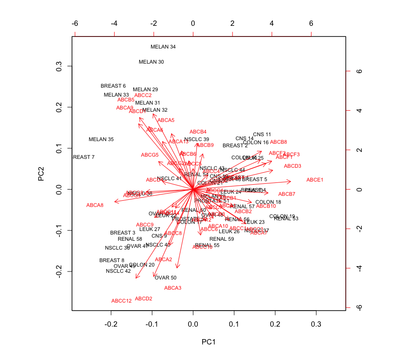
Multivariate methods for multiomics data analysis
|
A course dedicated to Multivariate data analysis for multi-omics data. Due to recent technological advancements, multi-omics has emerged as a trending approach to gain deeper insights into molecular mechanisms. Multi-omics approach involves integrating information from various level of central dogma of biology such as Genomics, Transcriptomics, Proteomics, Epigenetics and Metabolomics. Multi-omics has seen it's tremendous application in oncology. Since the information obtained at individual omics layer are highly heterogeneous, it is a challenging task to integrate them and derive useful information. This course aim at introducing you to various multivariate, clustering and integrative methods such as PCA, MDS, PLS, PLSDA, KC, HC, NMF etc., that are widely used in multi-omics data integration.
|
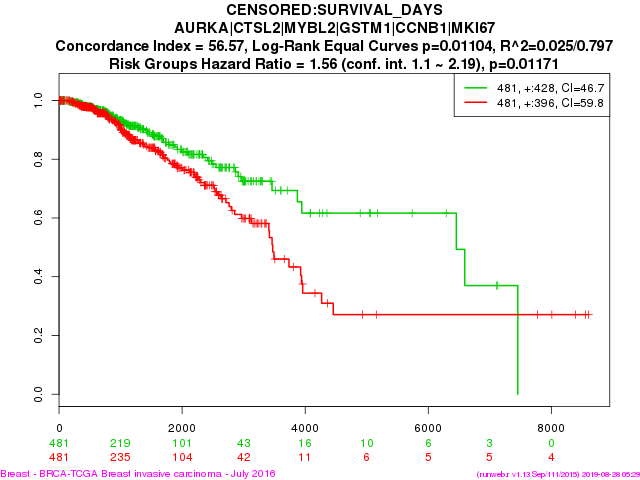
Survival Analysis using Omics Data
|
A course dedicated to survival data analysis using cancer omics data. Survival analysis is a widely used statistical method for studying time to event data. It has large scale applications in cancer.
|
Microarray Data Analysis
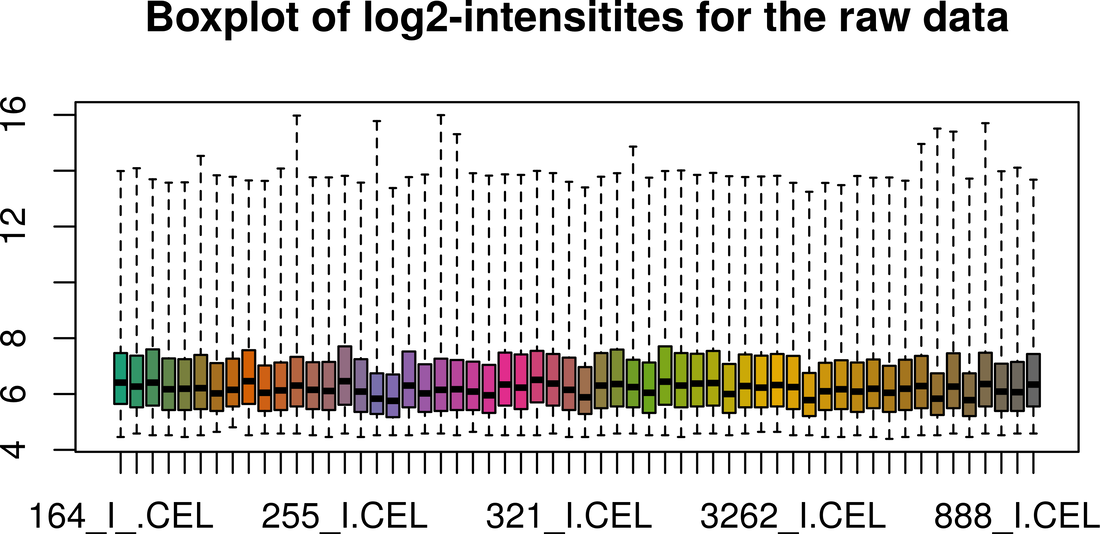
Gene Expression Analysis using Microarray
|
This course aims to illustrate the end to end workflow typically followed for analysis of Microarray based Gene Expression data. Although focuses on Affymetrix microarray data analysis, the steps can be applied to any other microarray platforms. Click on Know More to glance through course content.
|
Genome-Wide Human SNP Array 6.0, CNV analysis
|
This course aims to introduce you the steps followed for analysis of SNP 6.0 array data, specifically focused on deriving somatic copy-number alterations (SCNAs). The course discusses on CBS algorithm, tangent normalization method followed by running different tools for getting copy number information. The course ends with discussion on how TCGA used SNP 6.0 to obtain CNV data, their data processing steps. Click on Know More to glance through course content.
|
Methylation Array Data Analysis
|
This course is designed for analysis of Methylation array data. The course begins by introduction to micro array followed by different methylation platforms that are widely used. Click on Know More to glance through course content.
|
Introduction to Linux & Shell scripting
|
The objective of this course is to introduce you to Linux and Shell scripting. This will help you to perform common tasks in Linux environment without need to write lengthy codes. During the course we will cover various aspects of Linux, starting from understanding Linux terminal, file systems, permissions, navigating across different directory structures, basic file operations such as copy and delete. Click on know more to glance at the course.
|
MongoDB
|
MongoDB is one of the most popular NoSQL open source database. This course aims to train you on MongoDB basics such as installation, concept of databases, collections, document structure, querying the database. After the course you would be able to store any kind of unstructured data in MongoDB and play with the data via MongoQuery. This is useful in the field of Bioinformatics apart from MySQL language. Therefore we recommend you to learn this course.
|